Challenged with such a wealth of genome sequence data, to efficiently process and analyze these data, to compare similar regions and conserved sites between the two sequences, to seek sequence homology structures, and to reveal biological heredity, variation, and evolution, etc., have become the main motivations for the research of sequence alignment algorithms.Īt present, most of the research on alignment algorithms focus on specific problems ( Isa et al., 2014 Cattaneo et al., 2015 Chattopadhyay et al., 2015 Huo et al., 2016) or specific algorithm optimization ( Farrar, 2007 Houtgast et al., 2017 Junid et al., 2017) in the field of sequence similarity analysis, but less on the whole problem domain, so it is difficult to get an algorithm component library with a higher level of abstraction and suitable for the whole field of sequence similarity analysis. For example, Illumina HiSeqX Ten can generate approximately 3 billion 2 × 150 bp paired-end sequencing data within 3 days ( Illumina, 2016). 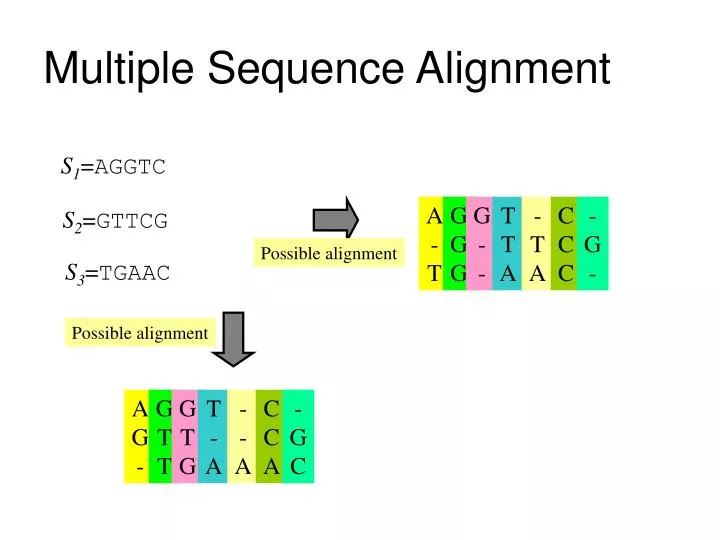
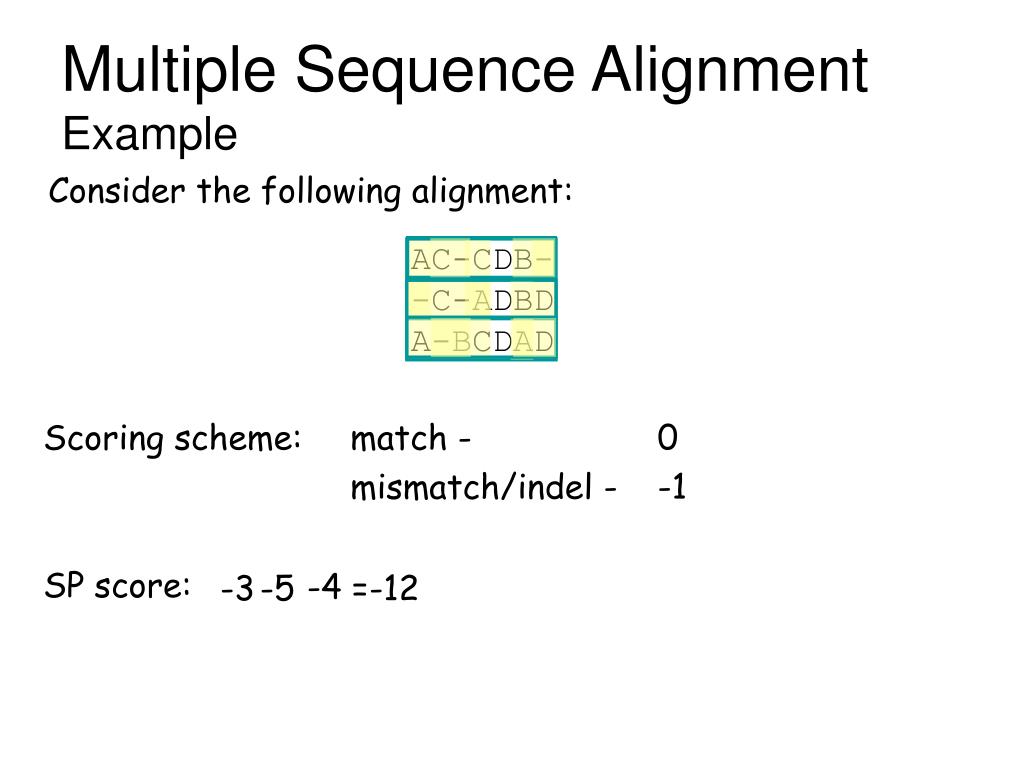
With the implementation of the Human Genome Project, the development of sequencing technology has produced a large amount of raw sequence data about biological molecules. Sequence alignment is a technique for identifying regions of sequence similarity by arranging genome sequences to obtain the function, structure, or evolutionary relationship between the sequences to be aligned. It is the key link to apply high-performance computing to biology. In the research of bioinformatics ( Wang et al., 2015), biological sequence alignment is one of the important processes of similarity analysis between unknown and known molecular sequences, the basis of biological sequence analysis and database search, and used in the sequence assembly. A star alignment algorithm is designed and generated to demonstrate the development process.Īlignment is a common and important approach in biology study. Based on our constructed pairwise sequence alignment algorithm component library and the convenient software platform PAR, a few expansion domain components are developed for multiple sequence alignment application domain, and specific multiple sequence alignment algorithm can be designed, and its corresponding program, i.e., C++/Java/Python program, can be generated efficiently and thus enables the improvement of the development efficiency of complex algorithms, as well as accuracy of sequence alignment calculation. Multiple sequence alignment algorithms are more complex, redundant, and difficult to understand, and it is not easy for users to select the appropriate algorithm some computing errors may occur. Existing research focuses mainly on the specific steps of the algorithm or is for specific problems, lack of high-level abstract domain algorithm framework. 3School of Computer and Information Engineering, Jiangxi Normal University, Nanchang, ChinaĪs a key algorithm in bioinformatics, sequence alignment algorithm is widely used in sequence similarity analysis and genome sequence database search.2School of Software, Jiangxi Normal University, Nanchang, China.1School of Information Management, Jiangxi University of Finance and Economics, Nanchang, China.Haipeng Shi 1,2, Haihe Shi 3 * and Shenghua Xu 1 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
